Homepage of the Biological Soft Matter Platform
Homepage of the “Momentum” Biomolecular Self-Assembly Research Group
Membrane Active Foldamers
Development of resistance by bacteria to antibiotics makes design of novel antimicrobial compounds increasingly important. As persistent cells often become slow-growing or dormant, strategies targeting their membrane are becoming more relevant. Toxic oligomers may assemble into hydrophilic or lipophilic sheet rich barrel constructs. However, this mechanism is not understood, greatly hindering rational development of similar compounds.
To approach this problem, we aim to design and study foldamer oligomer assemblies. We hope to define the fundamentals of how peptide – lipid bilayer interactions govern formation of potentially toxic oligomers at a molecular level. This may be exploited for developing new antimicrobial compounds.
Foldamers are highly similar to natural peptides in terms of structural diversity, thus they are ideal model systems, in several cases showing antibiotic activity and enzyme resistance. The structures will be designed with quantum mechanical (QM) and molecular dynamics MD tools and studied with experimental methods in model membranes. We use sum frequency generated spectroscopy (SFG), polarized light spectroscopy (CD, LD), X-ray scattering methods (SAXS, WAXS and X-ray reflectometry).
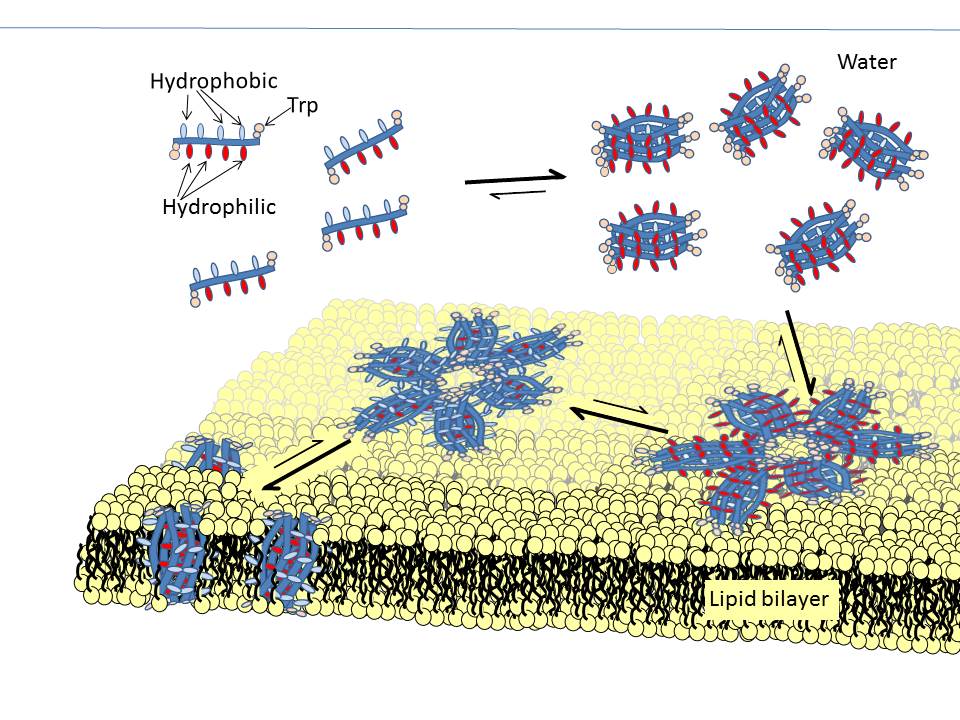
Figure 1. Proposed mechanism of oligomeric self-assembly into barrel scaffolds with environment-sensitive alternating side chain positions.
Related References
1) T. Keszthelyi, K. Hill, É Kiss Interaction of Phospholipid Langmuir Monolayers with an Antibiotic Peptide Conjugate J. Phys. Chem. B, 2013, 117 (23), pp 6969–6979
2) J. Johansson, E. Hermansson, N. Kann, B. Nordén, T. Beke-Somfai: d-Peptides from RuAAC-Derived 1,5-Disubstituted Triazole Units, Eur. J. Org. Chem. 2014, 13, 2703-2713
3) N. Kann, J. R. Johansson, T. Beke-Somfai*: Conformational properties of 1,4- and 1,5-substituted 1,2,3-triazole amino acids – building units for peptidic foldamers, Organic & Biomolecular Chemistry, 2015, DOI: 10.1039/C4OB02359E
4) T. Beke, A. Czajlik, B. Bálint, A. Perczel: A Theoretical Comparison of Self-Assembling a- and b-Peptide Nanostructures: Toward Design of b-Barrel Frameworks ACS Nano, 2008, 2, 545-553
5) T. Beke, I. G. Csizmadia, A. Perczel: Theoretical study on tertiary structural elements of b-peptides: Nanotubes formed from parallel-sheet-derived assemblies of b-peptides J. Am. Chem. Soc., 2006, 128, 5158-5167
Complex Biomolecular Machines
In a wide range of biological activities, from cell locomotion to membrane transport, nature deploys numerous sophisticated molecular machines which have become highly optimized for performance and controllability. Rational design and engineering of similarly complex biosystems is a very exciting field with a potential to dramatically alter future’s medicine or industrial biochemistry. However, to overcome major challenges in design of artificial enzymes, the precise understanding of control mechanism on key reaction steps by larger molecular scale structure and dynamics is required. FoF1 ATP synthase is interesting as a model system: a delicate molecular machine synthesizing or hydrolysing ATP utilizing a rotary motor (1-2). ATP synthase is a member of the RecA-like helicase family, and it is particularly interesting how the structural and residual differences of the same family determine the ATP hydrolysis mechanism and its effect on the overall function of these enzymes. Rad51, RadA, and RecA are examples from this group of proteins, which fulfil particularly important roles in cellular functions: the repair of damaged DNA and the maintenance of genomic diversity. Despite the numerous studies, many details are still uncovered, especially the role of ATP hydrolysis and the description of the reaction mechanism at the atomic level. Computational biophysics provides an adequate set of tools to describe the atomic level structural differences and accurate energetics of the system. By using our experience in different QM and (MD) simulations, we would like to expand our current understanding of RecA enzymes, aiming at providing details on the coupling of reaction steps with the large scale motions (3-5).
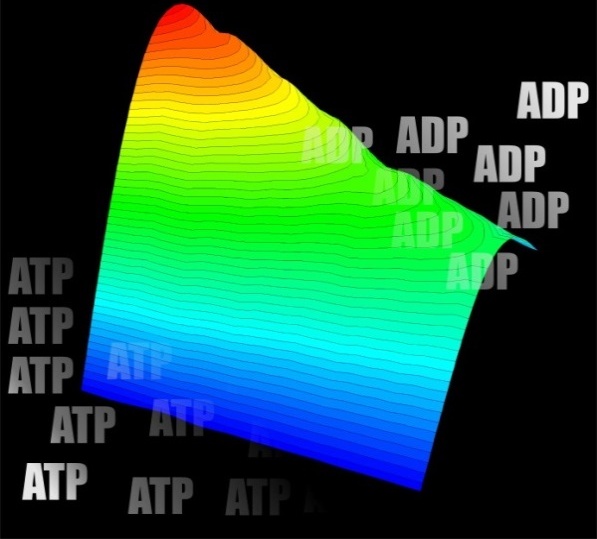
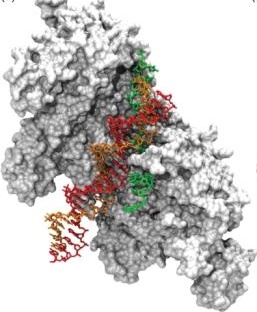
Figure 2. LEFT: Energy surface of ATP synthesis/hydrolysis in ATP synthase. RIGHT: The surface model of RecA filament with double-stranded DNA (image from: openi.nlm.nih.gov)
Related References
(1) T. Beke-Somfai et al. PNAS 2011, 108, 4828
(2) T. Beke-Somfai et al. PNAS 2013, 110, 2117-2122
(3) X. Qian, et al J. Biol. Chem. 2006, 281, 39380
(4) Z. Chen, et al Nature, 2008 453, 489
(5) A. Reymer et al. Biochemistry, 2015, 54, 4579
Leader
Tamás Beke-Somfai